Learning Objectives
Create mosaics of more than one multiband image
Find the intersection of two images
View the footprint of multiple image tiles
Handle Multiple Remotely Sensed Images#
Doing analysis over larger areas often requires the use of image mosaics (combining two or more images). Luckily for us geowombat makes this process relatively easy.
Union (Mosaic) of Remotely Sensed Image#
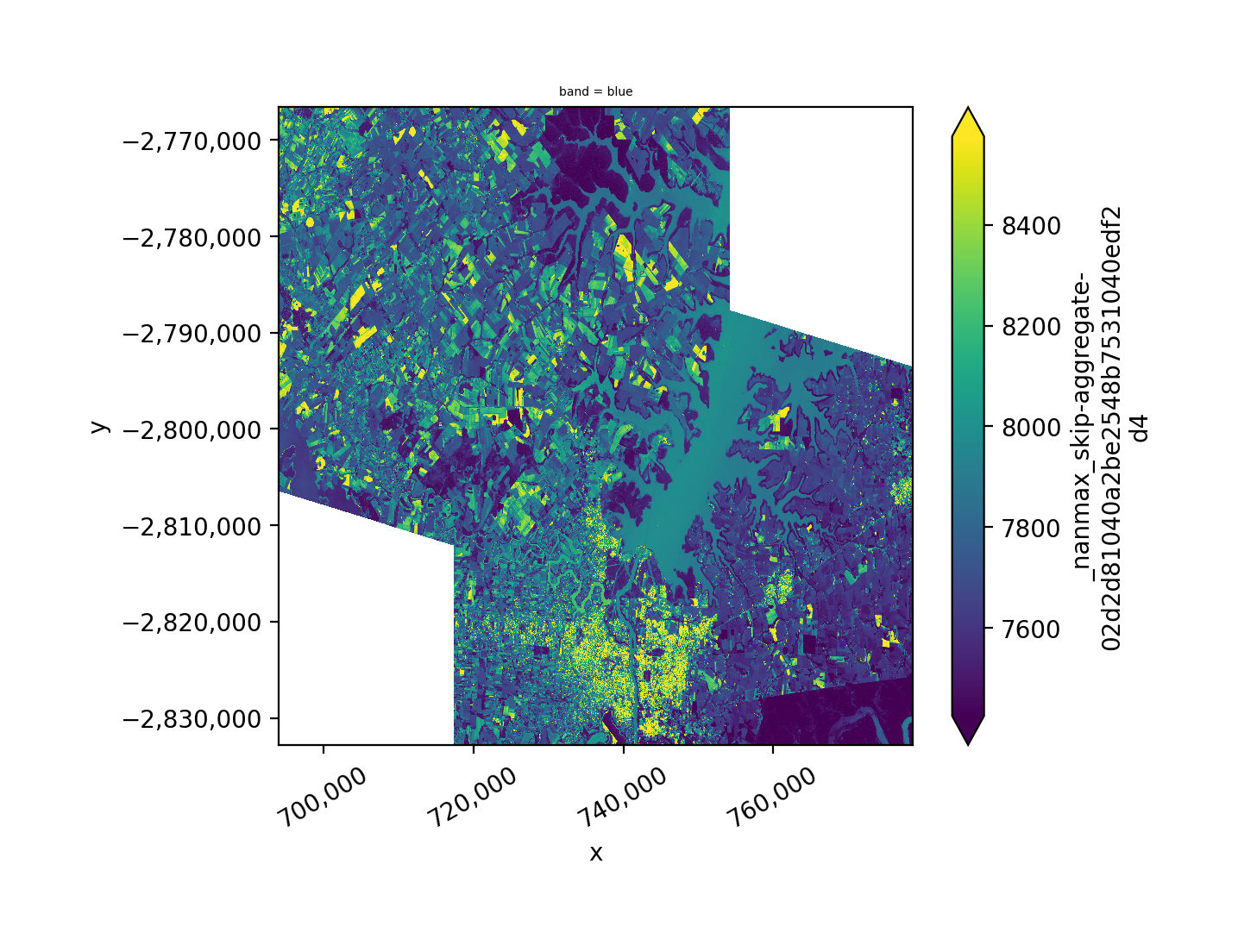
As an example let’s plot the union with mosaic=True of two images taken on the same day, for the overlapping portions we will use the mean pixel value by setting overlap='mean', but blue band only. Alternatively we could use one of ‘mean’, ‘min’, or ‘max’.
Note we rename the band name with band_names=['blue'].
# Import GeoWombat
import geowombat as gw
# import plotting
import matplotlib.pyplot as plt
import matplotlib.patheffects as pe
# load data
from geowombat.data import l8_224077_20200518_B2, l8_224078_20200518_B2
fig, ax = plt.subplots(dpi=200)
with gw.open(
[l8_224077_20200518_B2, l8_224078_20200518_B2],
band_names=['blue'],
mosaic=True,
bounds_by='union'
) as src:
src.where(src != 0).sel(band='blue').gw.imshow(robust=True, ax=ax)
plt.tight_layout(pad=1)
Intersection of Remotely Sensed Image#
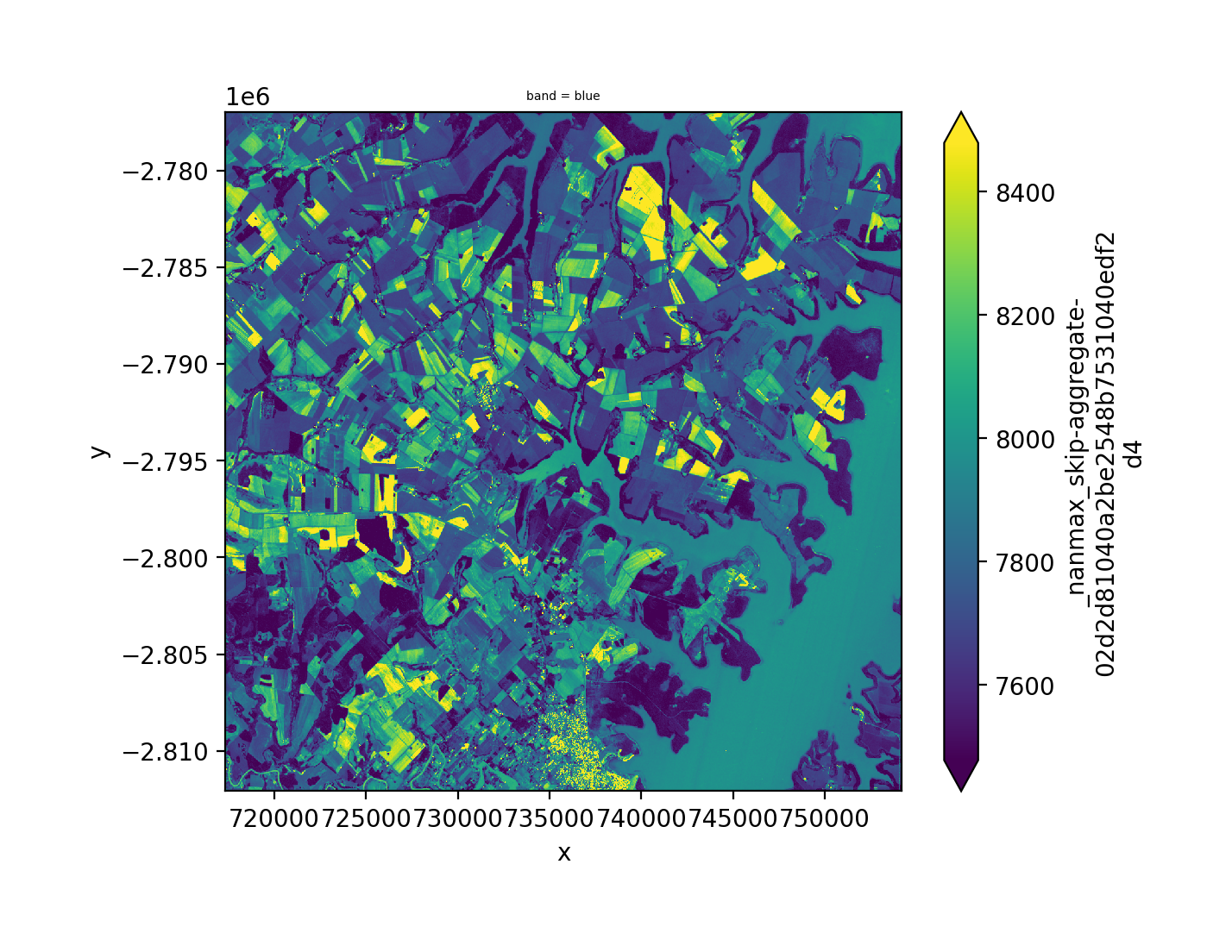
Same idea with the intersection, using bounds_by='intersection', we still need to mosaic the two images mosaic=True.
fig, ax = plt.subplots(dpi=200)
filenames = [l8_224077_20200518_B2, l8_224078_20200518_B2]
with gw.open(filenames,
band_names=['blue'],
mosaic=True,
overlap='max',
bounds_by='intersection') as src:
src.where(src != 0).sel(band='blue').plot.imshow(robust=True, ax=ax)
plt.tight_layout(pad=1)